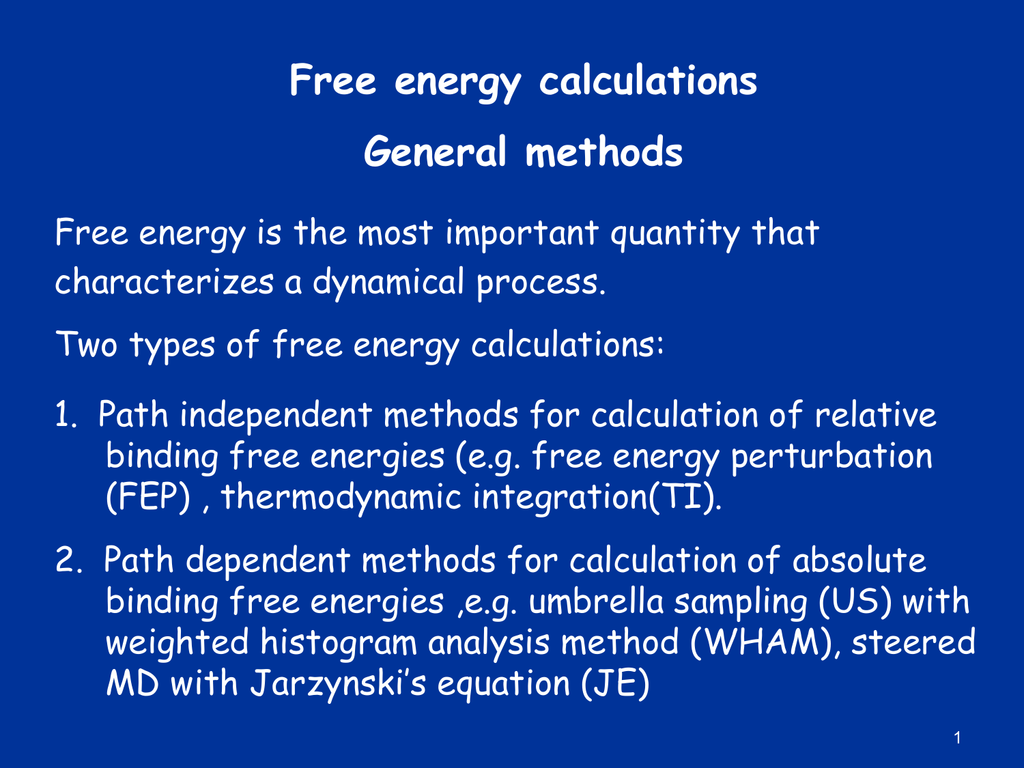
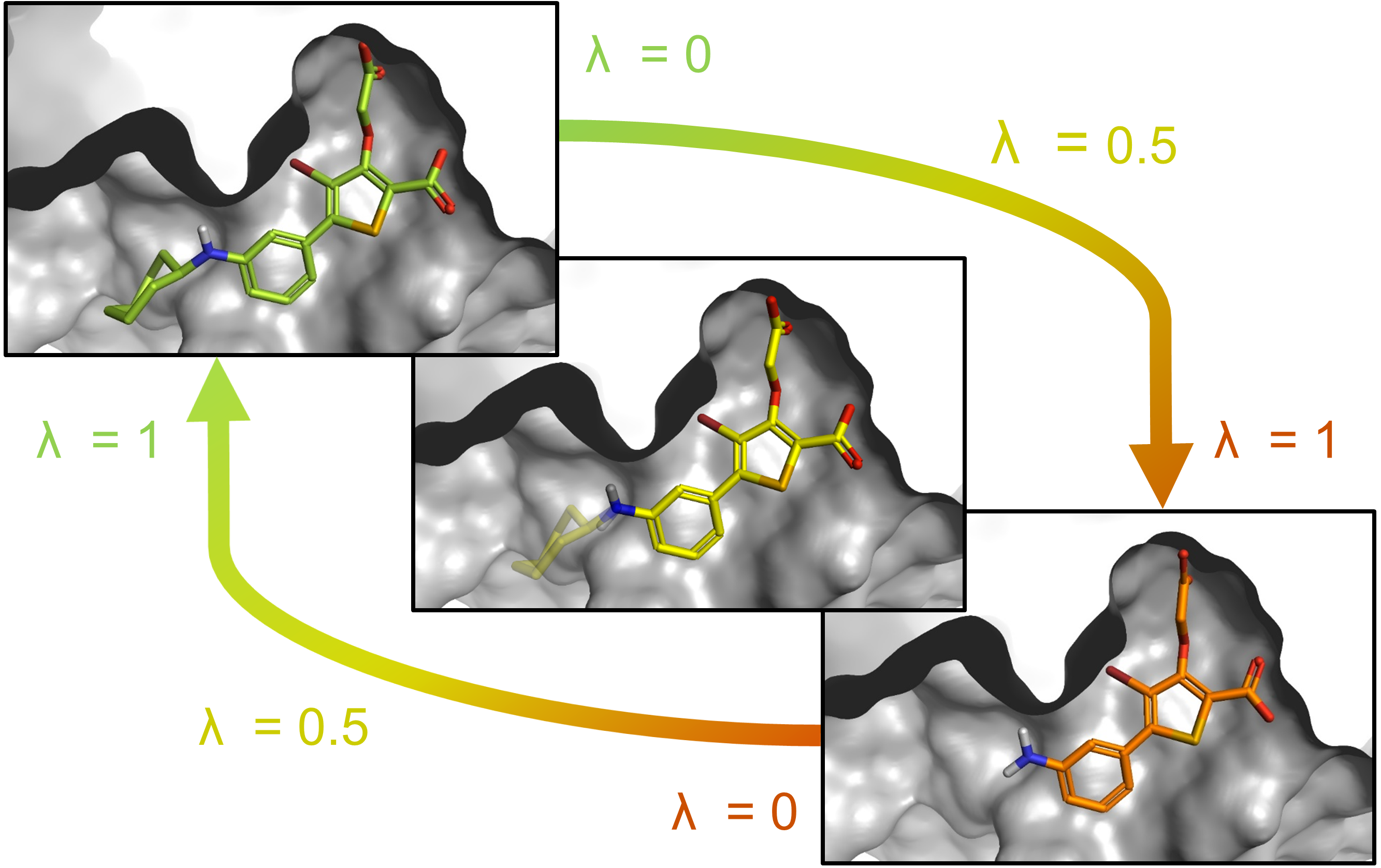
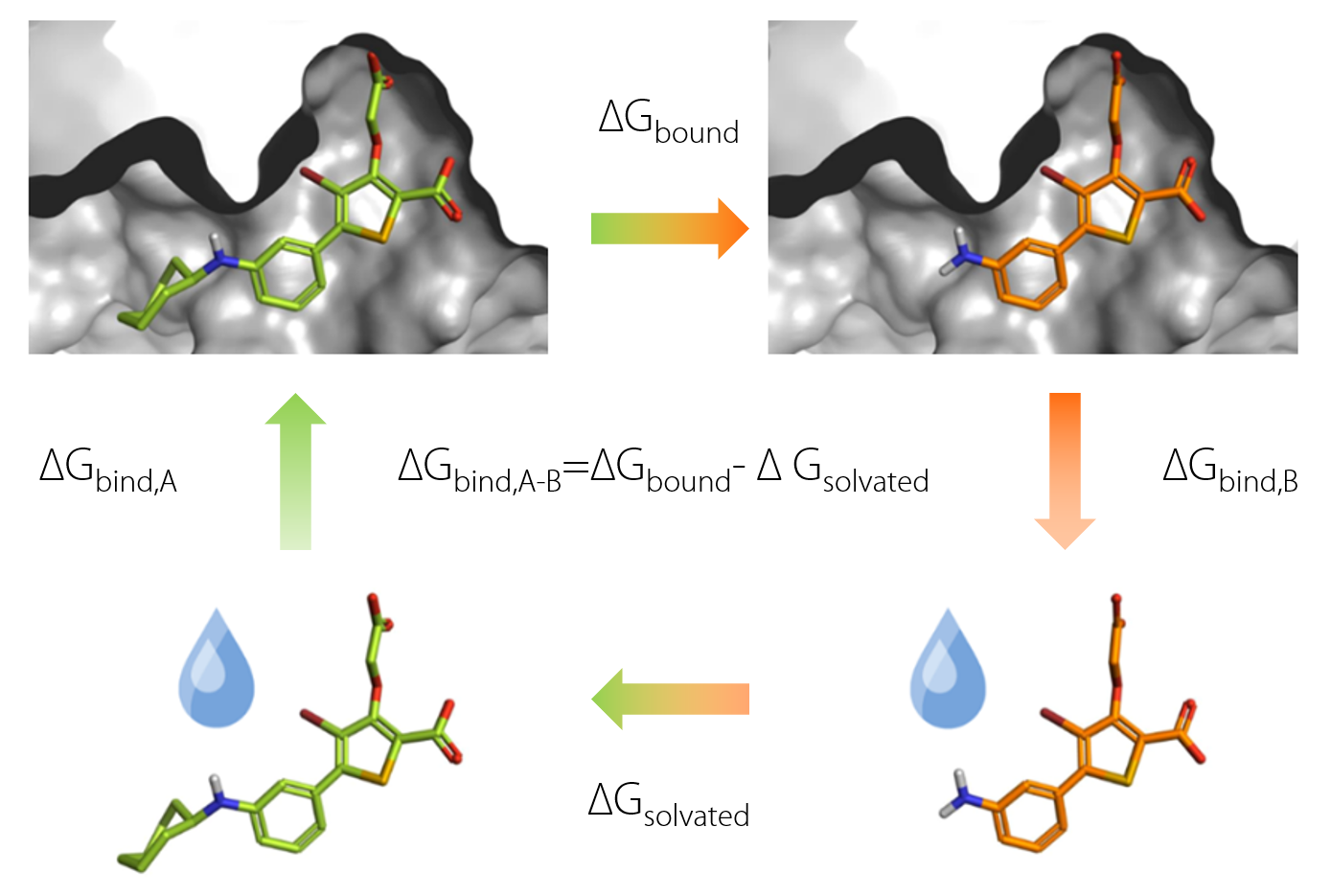
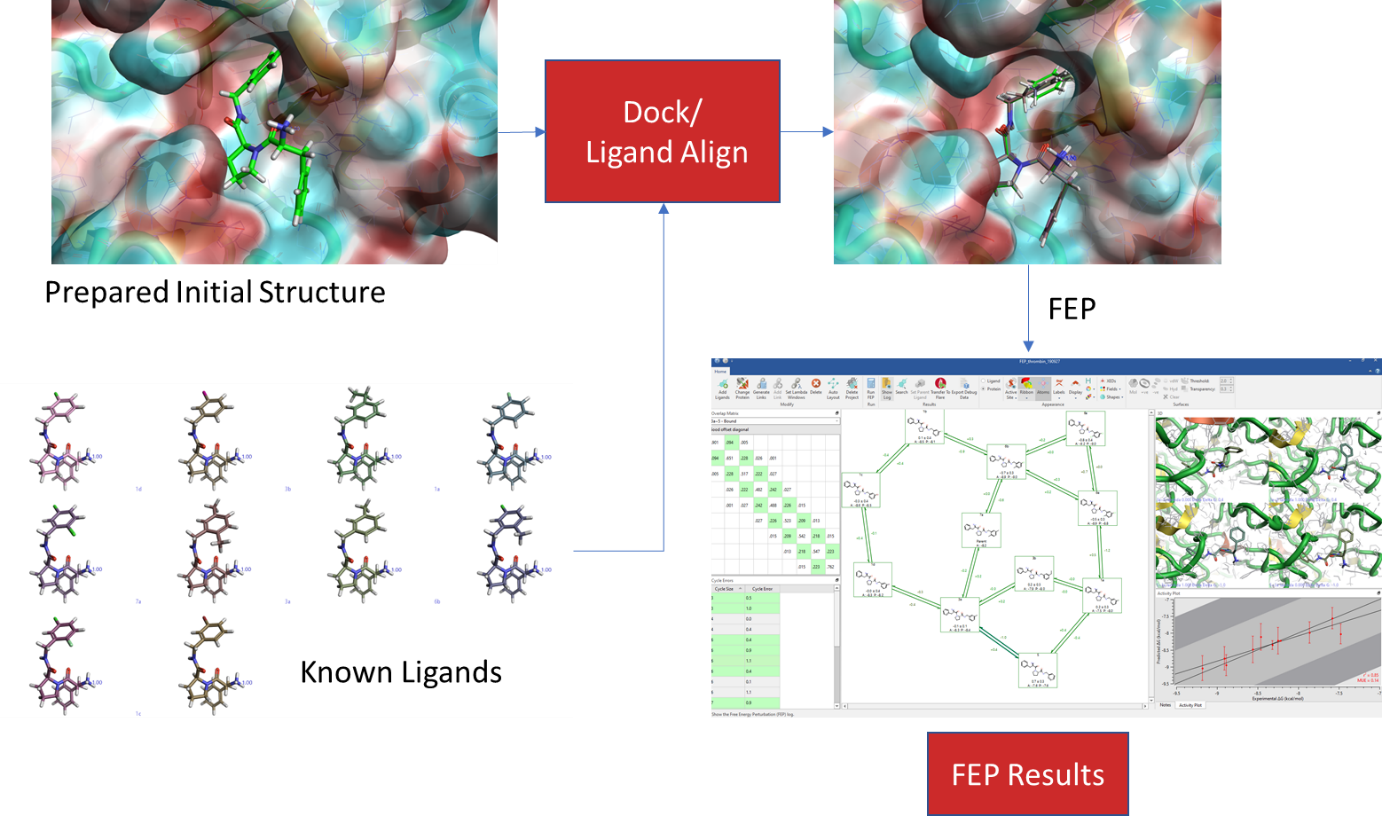
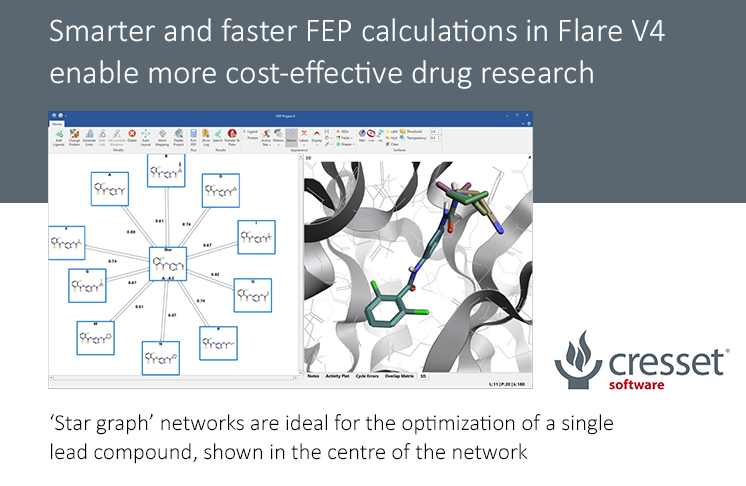
An example of FEP calculation on core-hopping using fragment-based design. | Download Scientific Diagram
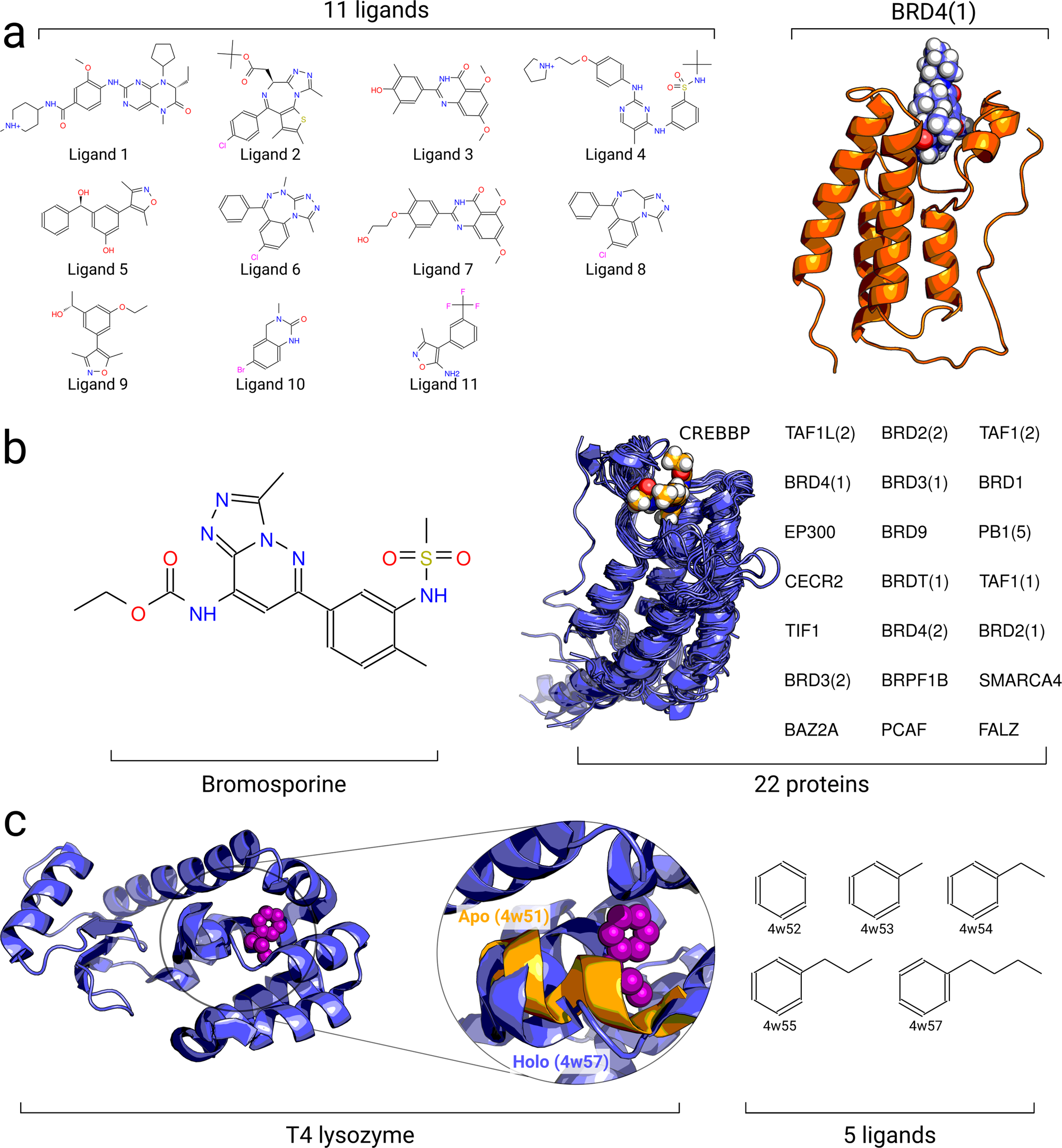
Accurate absolute free energies for ligand–protein binding based on non-equilibrium approaches | Communications Chemistry
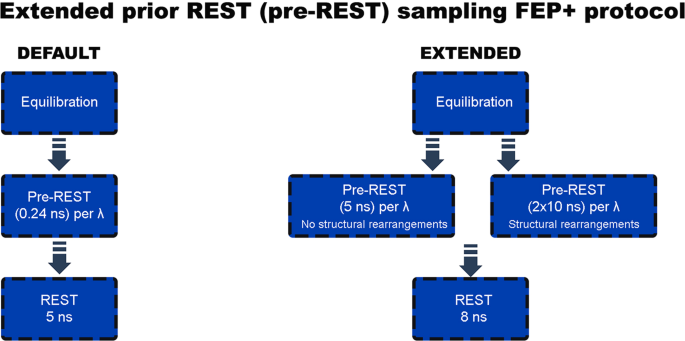
An Improved Free Energy Perturbation FEP+ Sampling Protocol for Flexible Ligand-Binding Domains | Scientific Reports

Relative Binding Affinity Prediction of Charge-Changing Sequence Mutations with FEP in Protein–Protein Interfaces - ScienceDirect

Modified Hamiltonian in FEP Calculations for Reducing the Computational Cost of Electrostatic Interactions | Journal of Chemical Information and Modeling

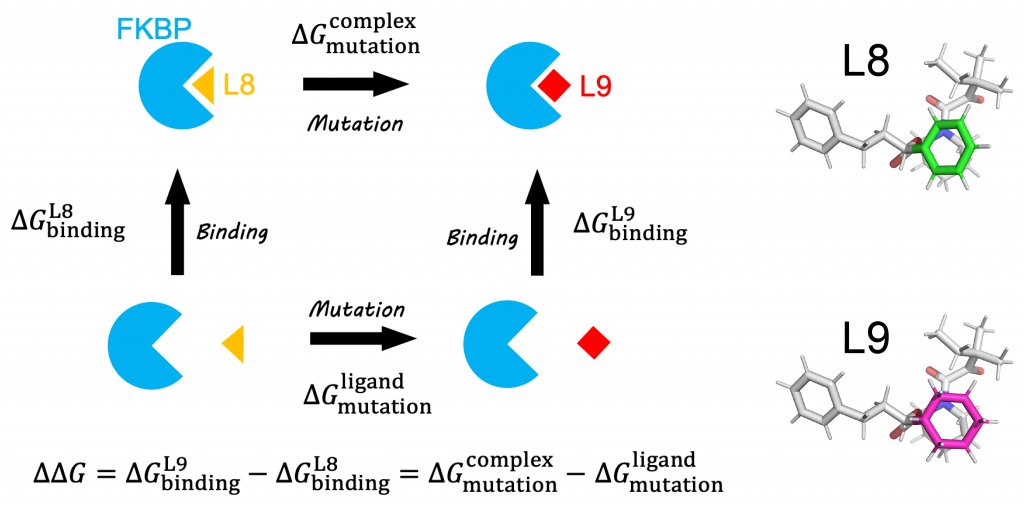
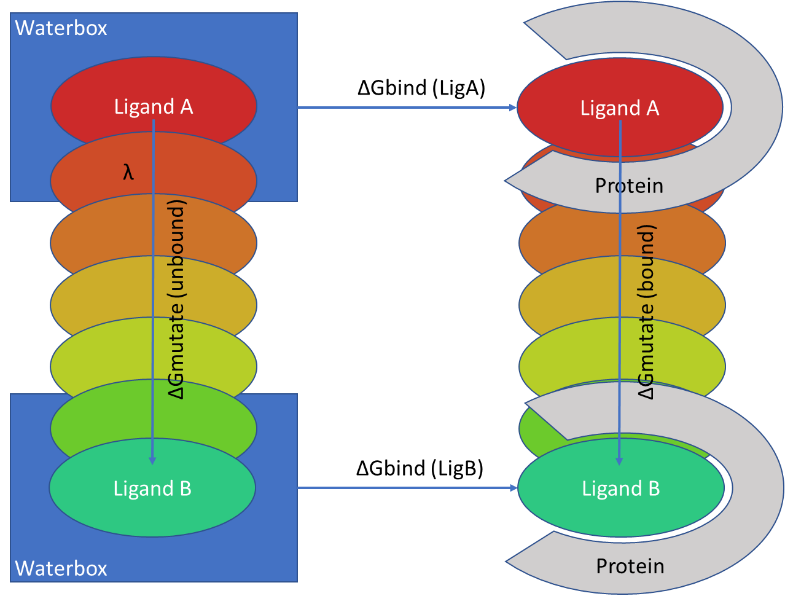
Description of the FEP calculations. A , thermodynamic cycle for FEP... | Download Scientific Diagram

FEP calculation of the relative binding free energy due to a mutation.... | Download Scientific Diagram
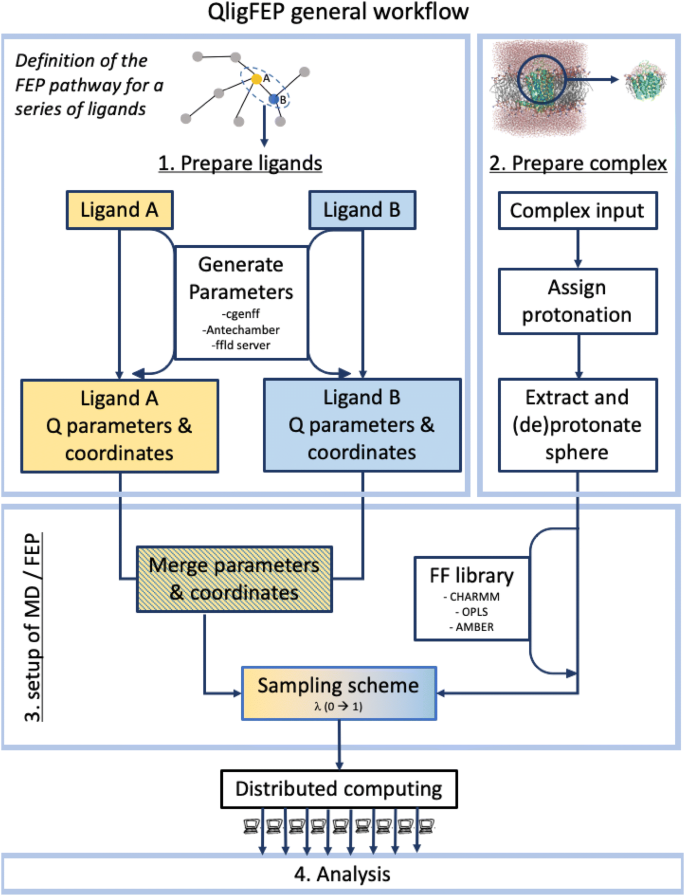
QligFEP: an automated workflow for small molecule free energy calculations in Q | Journal of Cheminformatics | Full Text
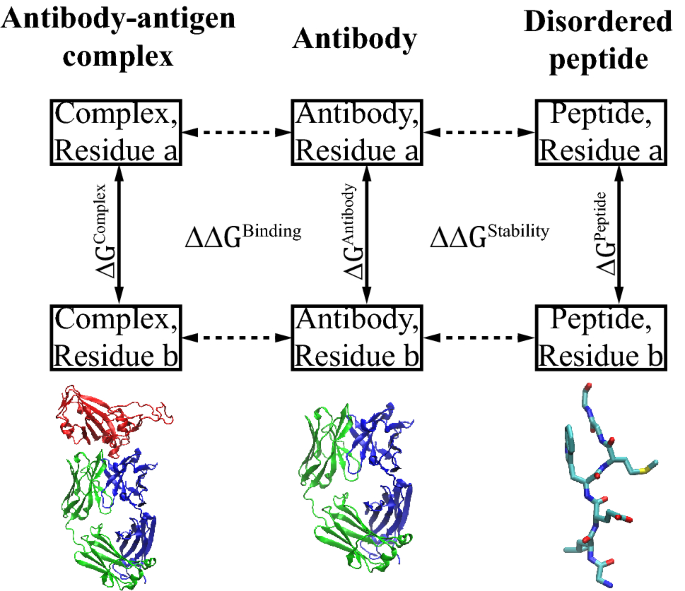
Large-scale application of free energy perturbation calculations for antibody design | Scientific Reports