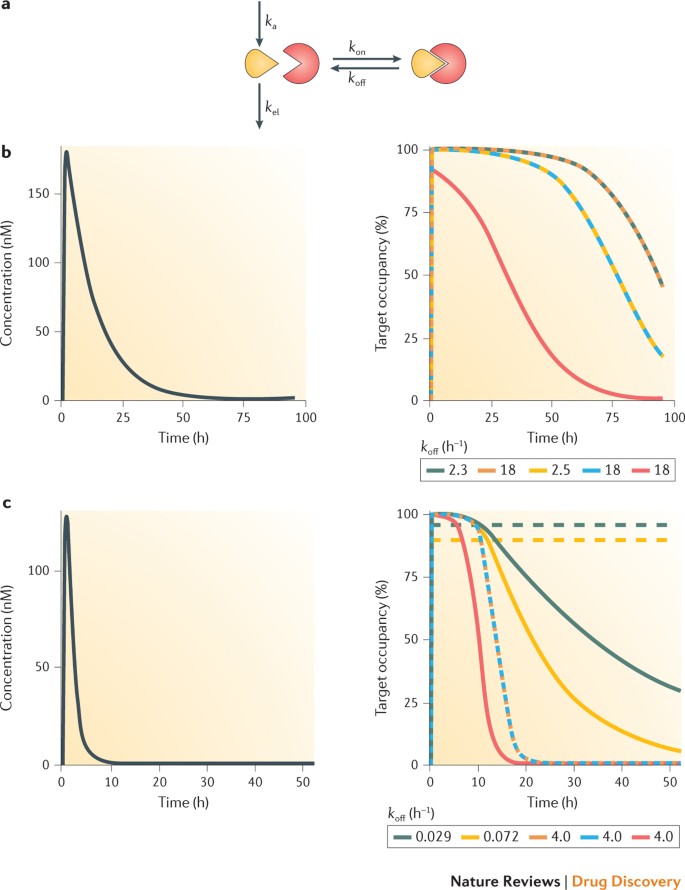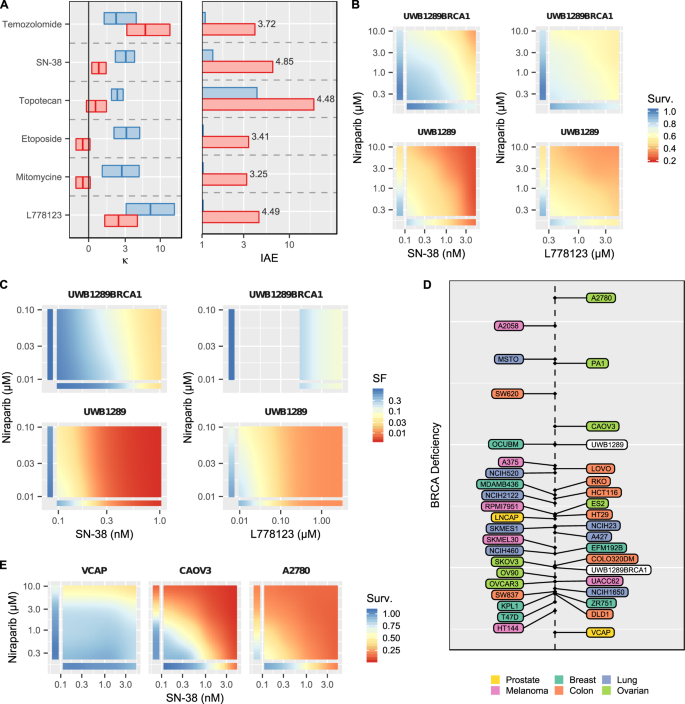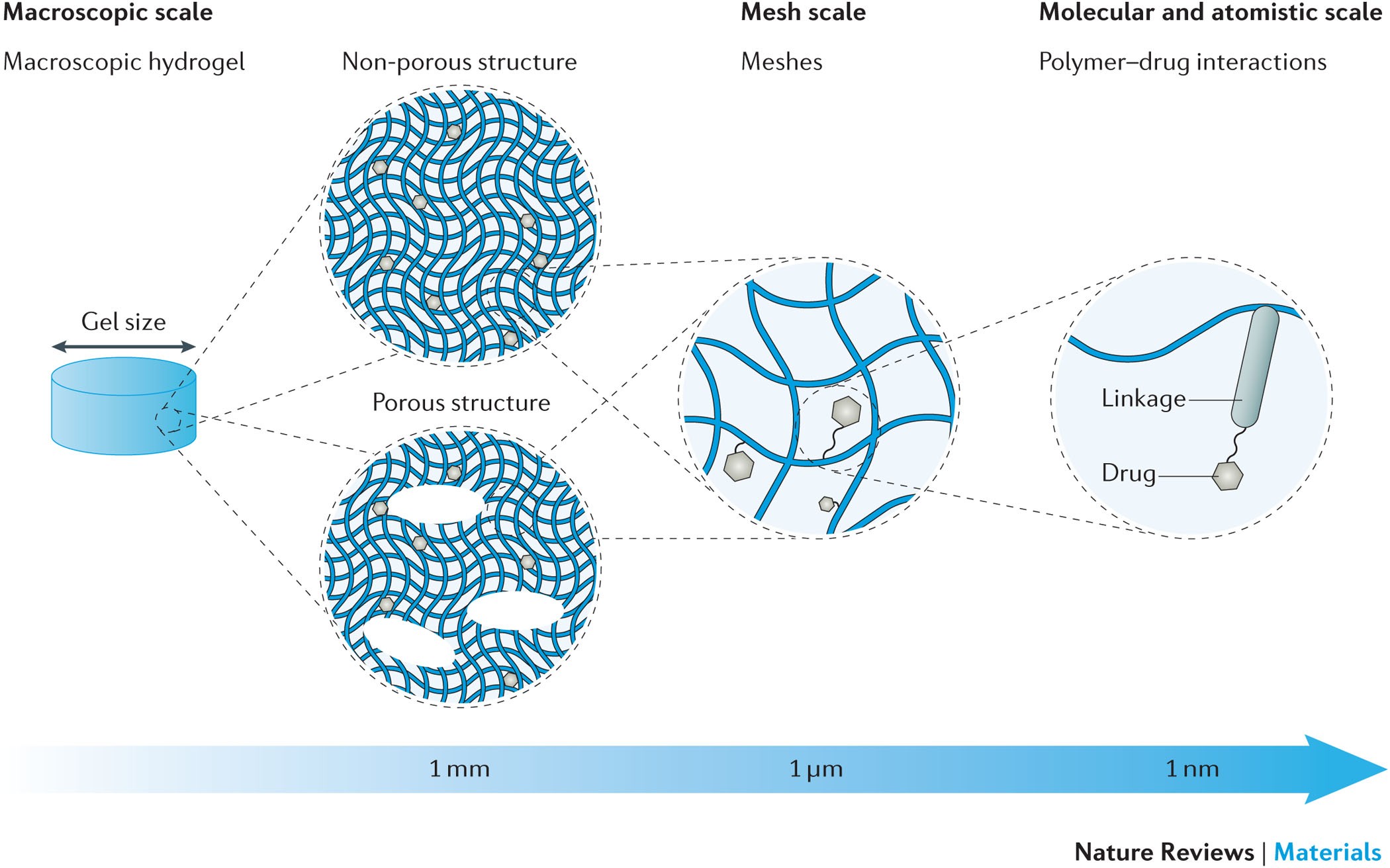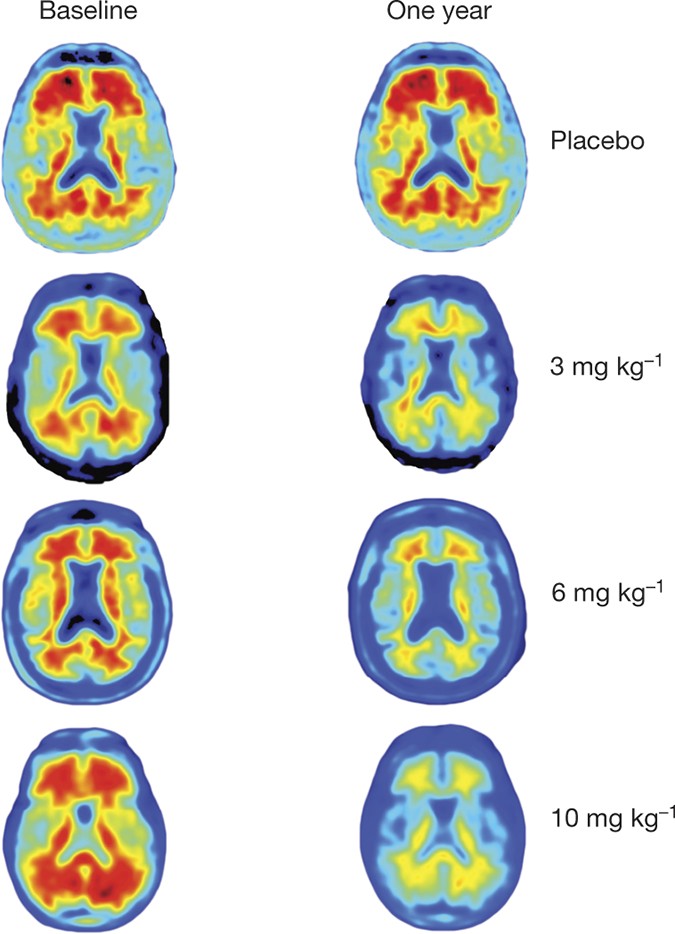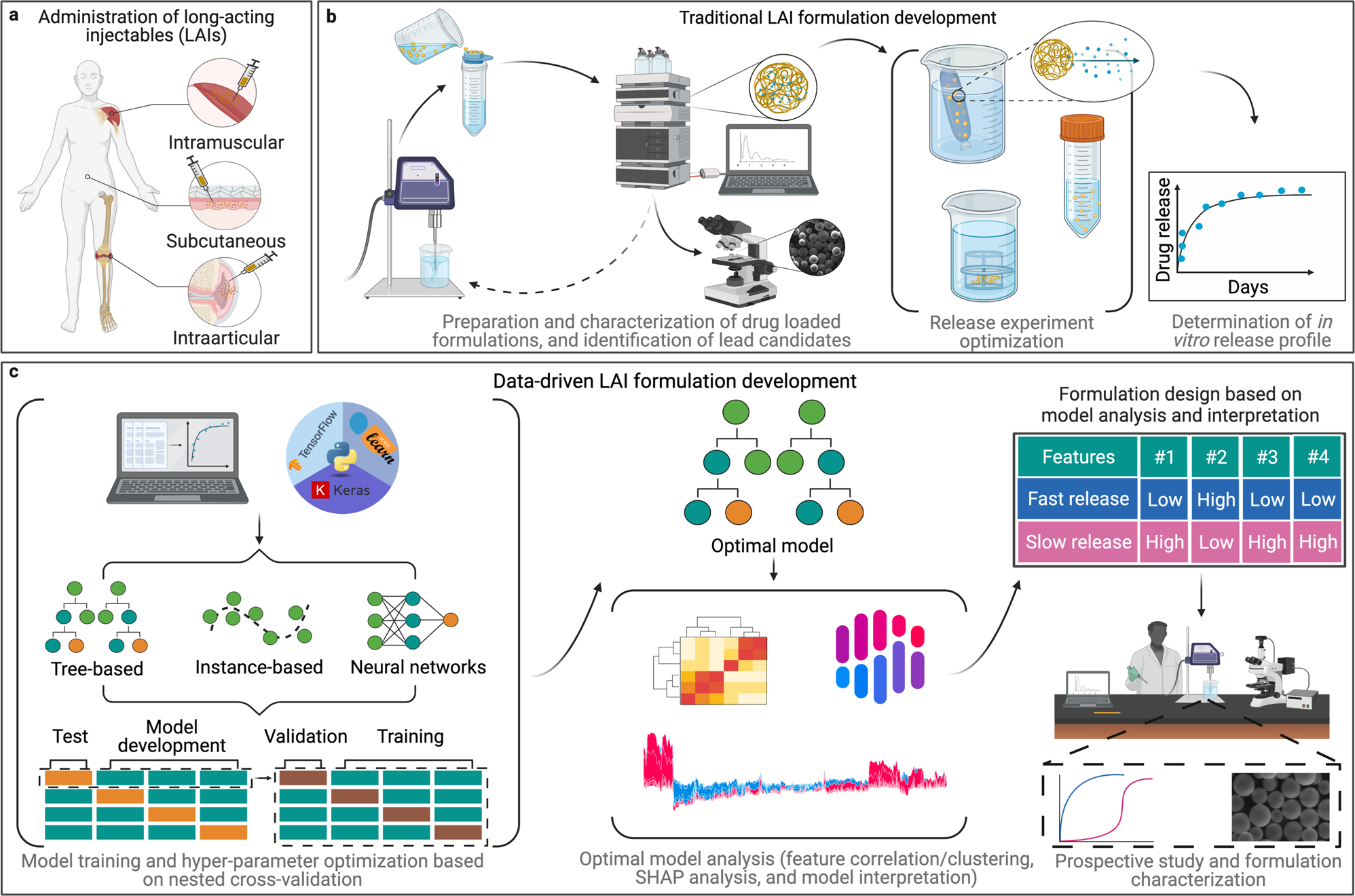
Machine learning models to accelerate the design of polymeric long-acting injectables | Nature Communications
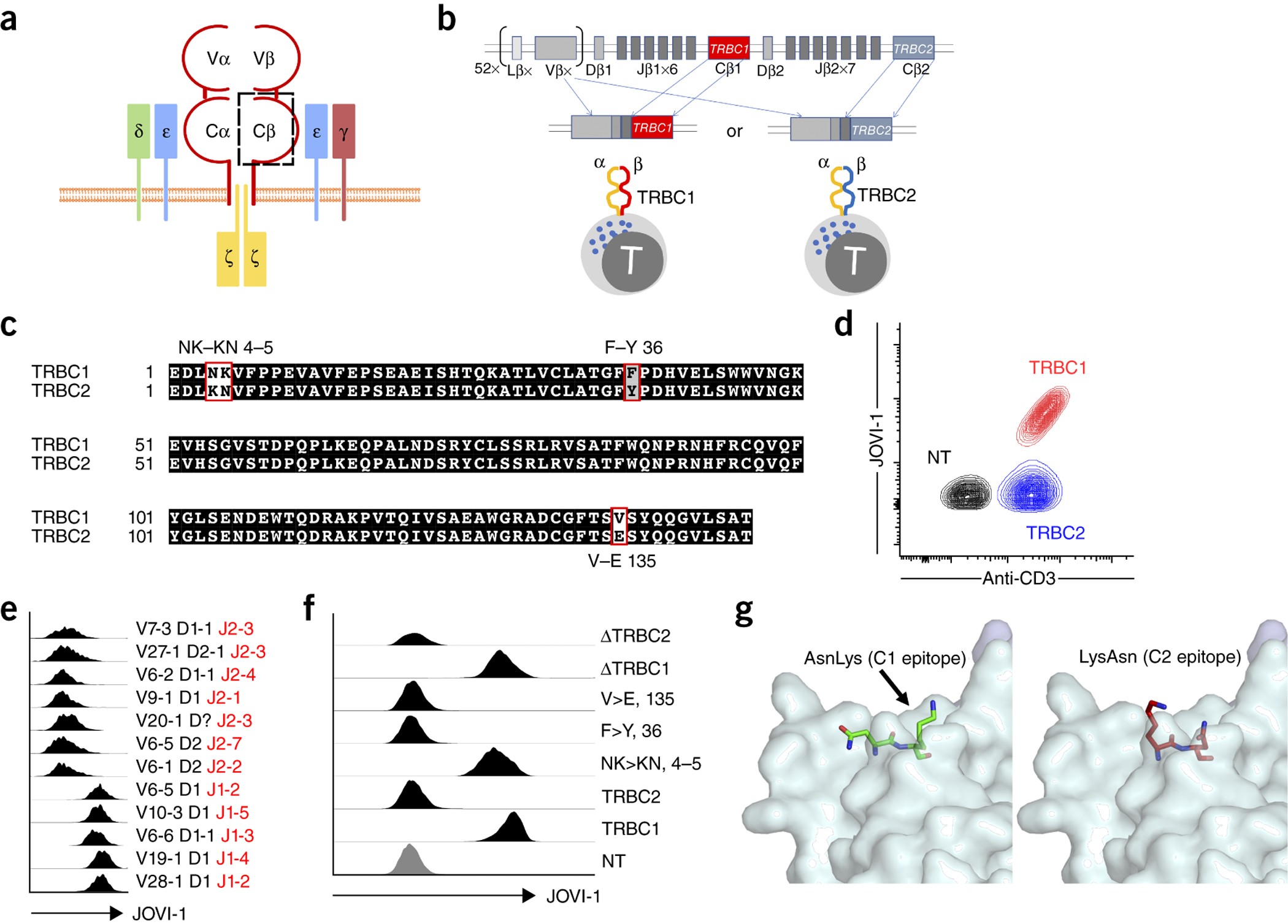
Targeting the T cell receptor β-chain constant region for immunotherapy of T cell malignancies | Nature Medicine
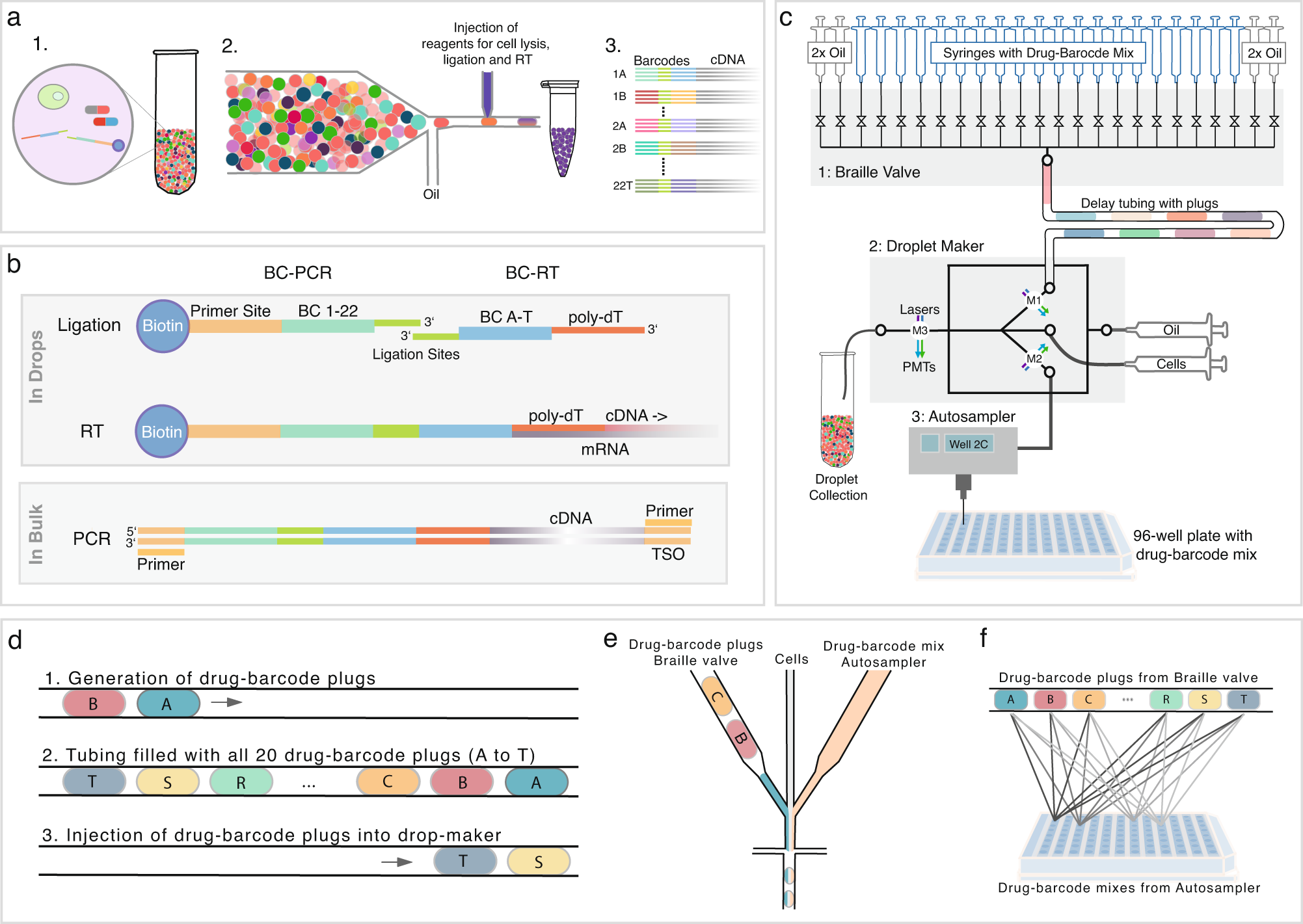
Combi-seq for multiplexed transcriptome-based profiling of drug combinations using deterministic barcoding in single-cell droplets | Nature Communications
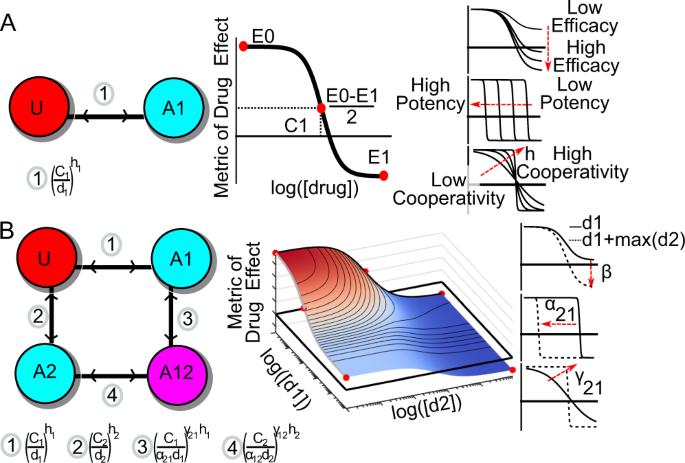
MuSyC is a consensus framework that unifies multi-drug synergy metrics for combinatorial drug discovery | Nature Communications

Synergistic drug combinations tend to improve therapeutically relevant selectivity | Nature Biotechnology
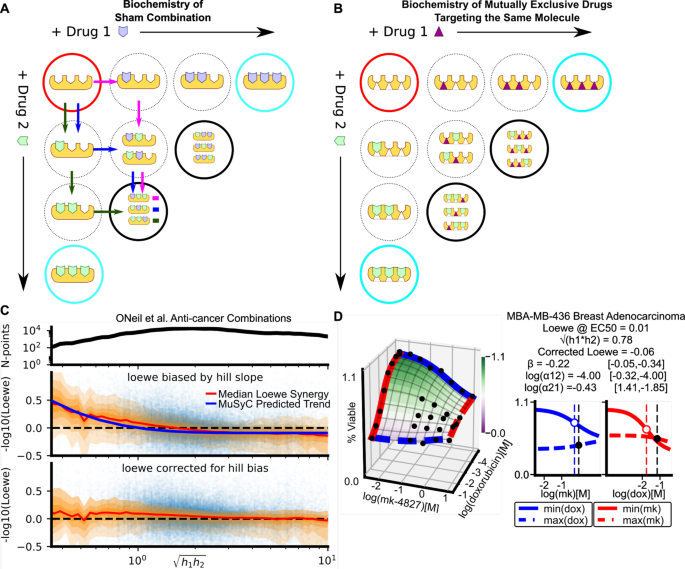
MuSyC is a consensus framework that unifies multi-drug synergy metrics for combinatorial drug discovery | Nature Communications
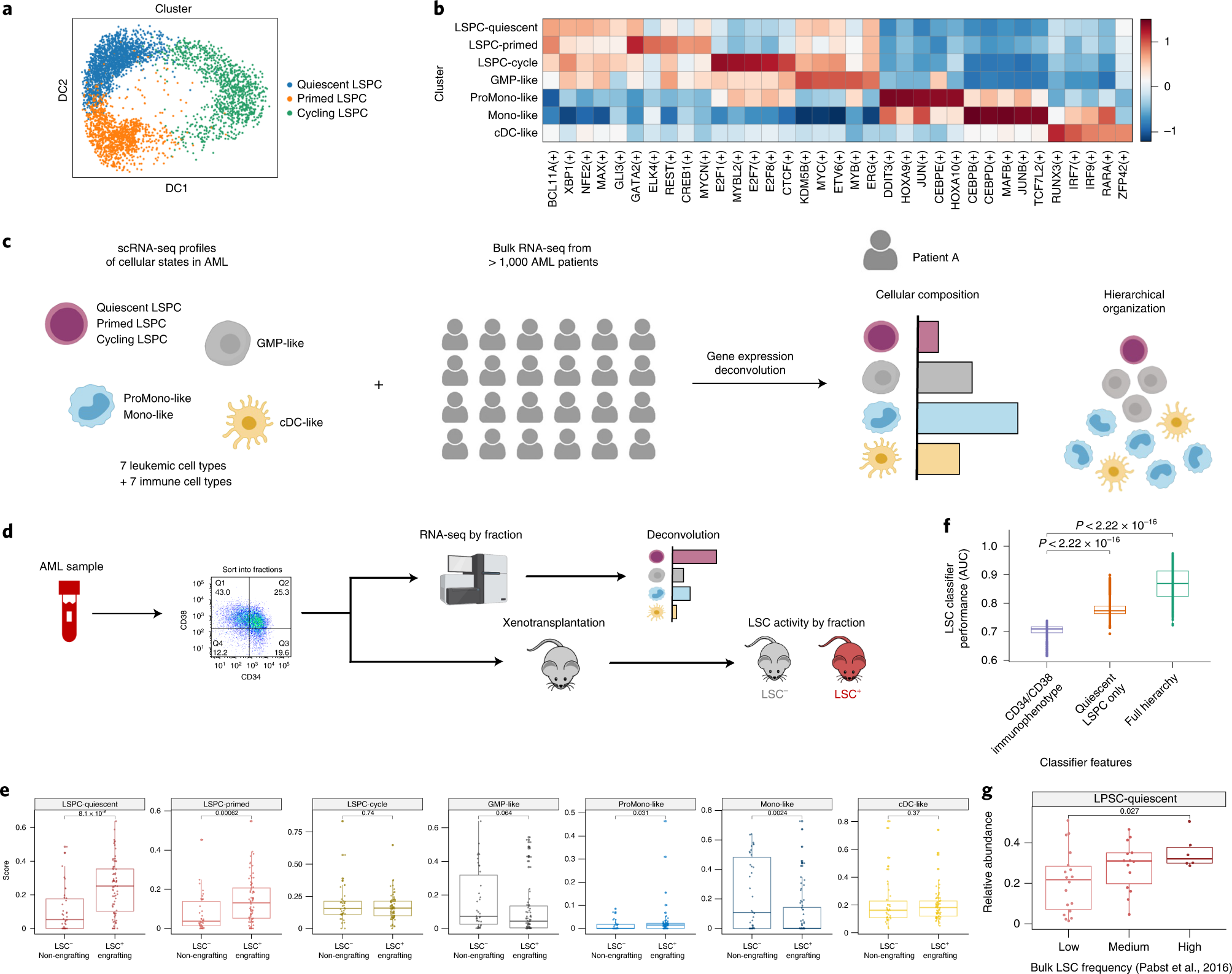
A cellular hierarchy framework for understanding heterogeneity and predicting drug response in acute myeloid leukemia | Nature Medicine

A machine learning-based chemoproteomic approach to identify drug targets and binding sites in complex proteomes | Nature Communications
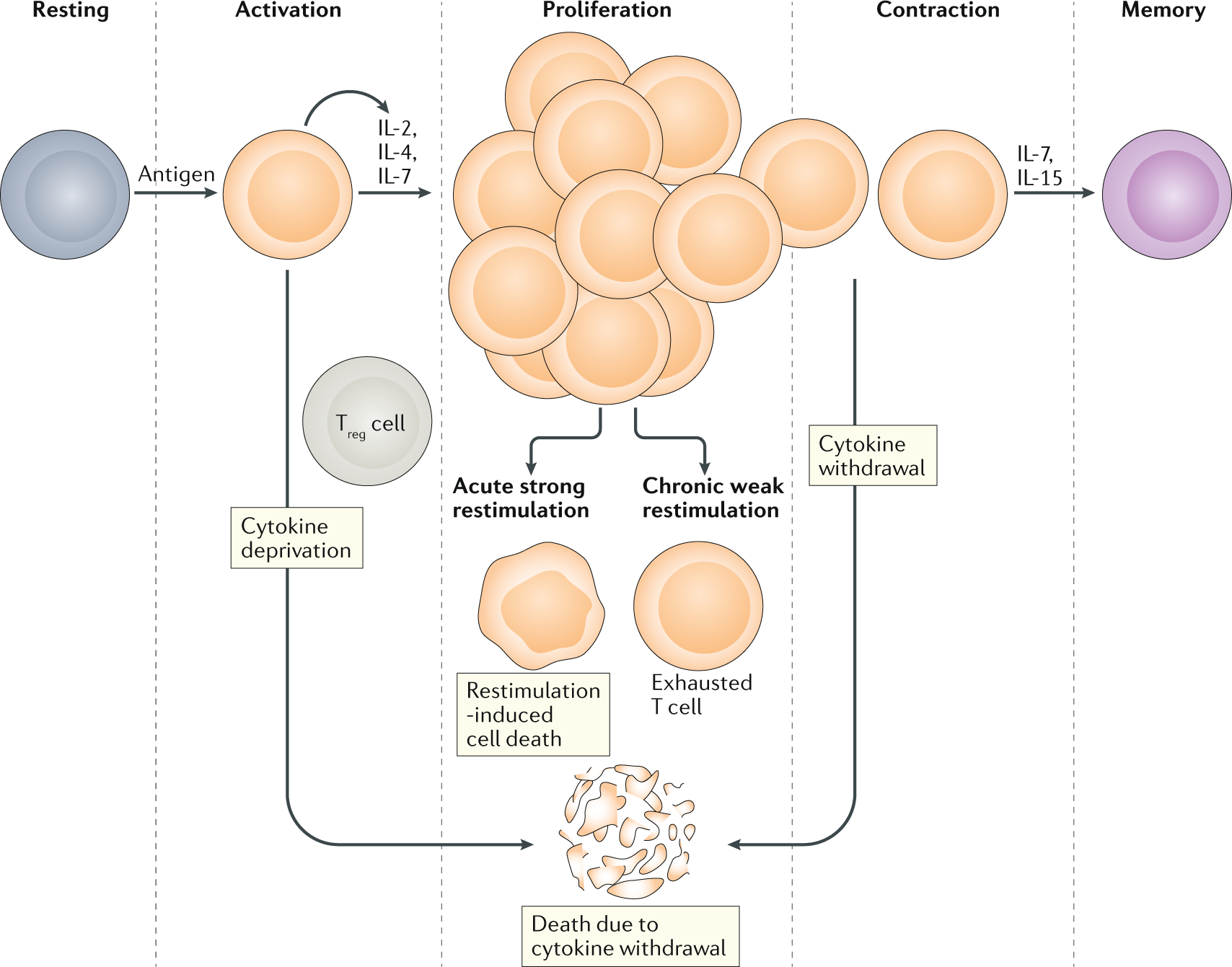
A guide to cancer immunotherapy: from T cell basic science to clinical practice | Nature Reviews Immunology

DNA-encoded chemistry: enabling the deeper sampling of chemical space | Nature Reviews Drug Discovery

Determination of the ratio of drug combinations to induce synergy. (A,... | Download Scientific Diagram
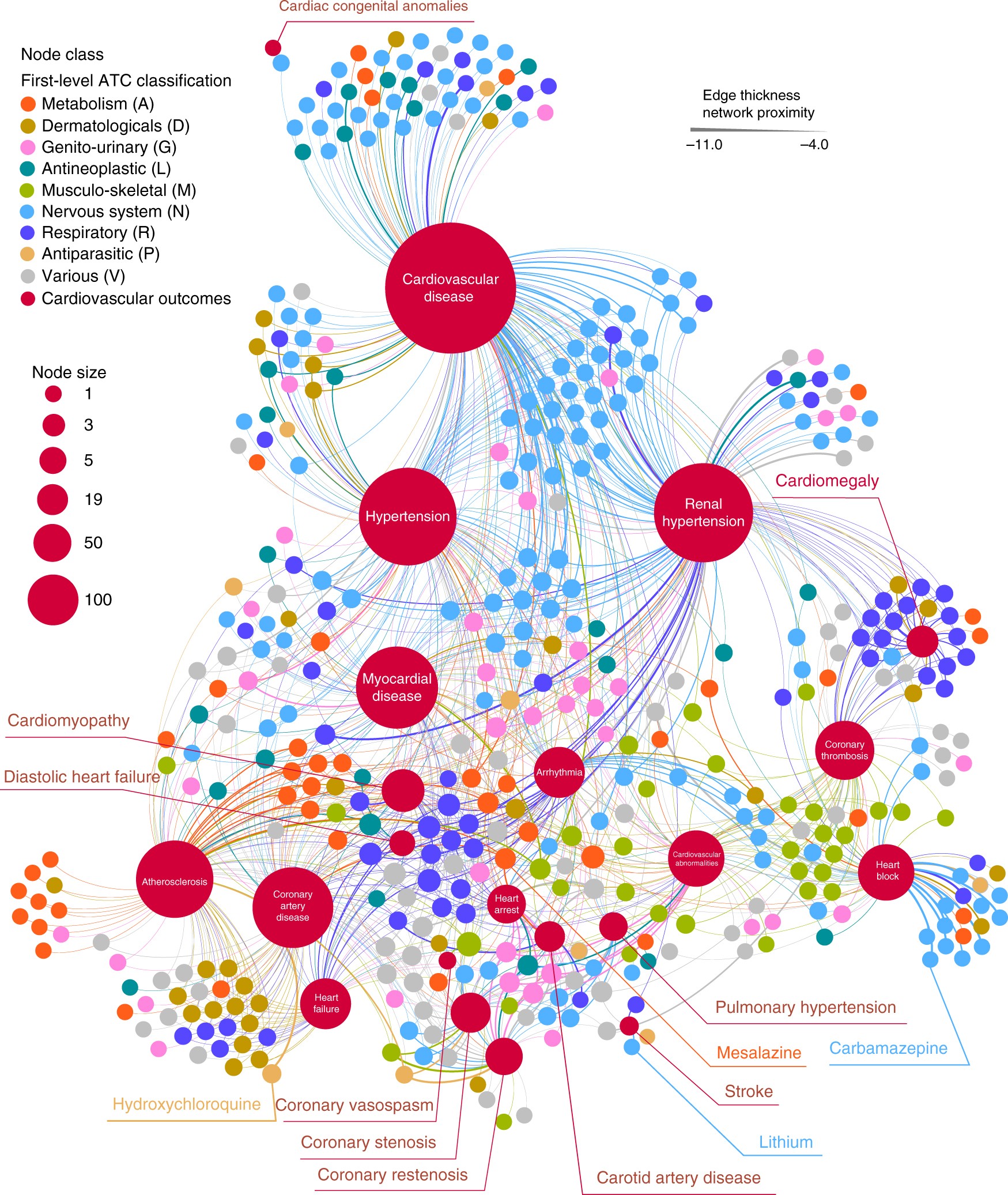
Network-based approach to prediction and population-based validation of in silico drug repurposing | Nature Communications
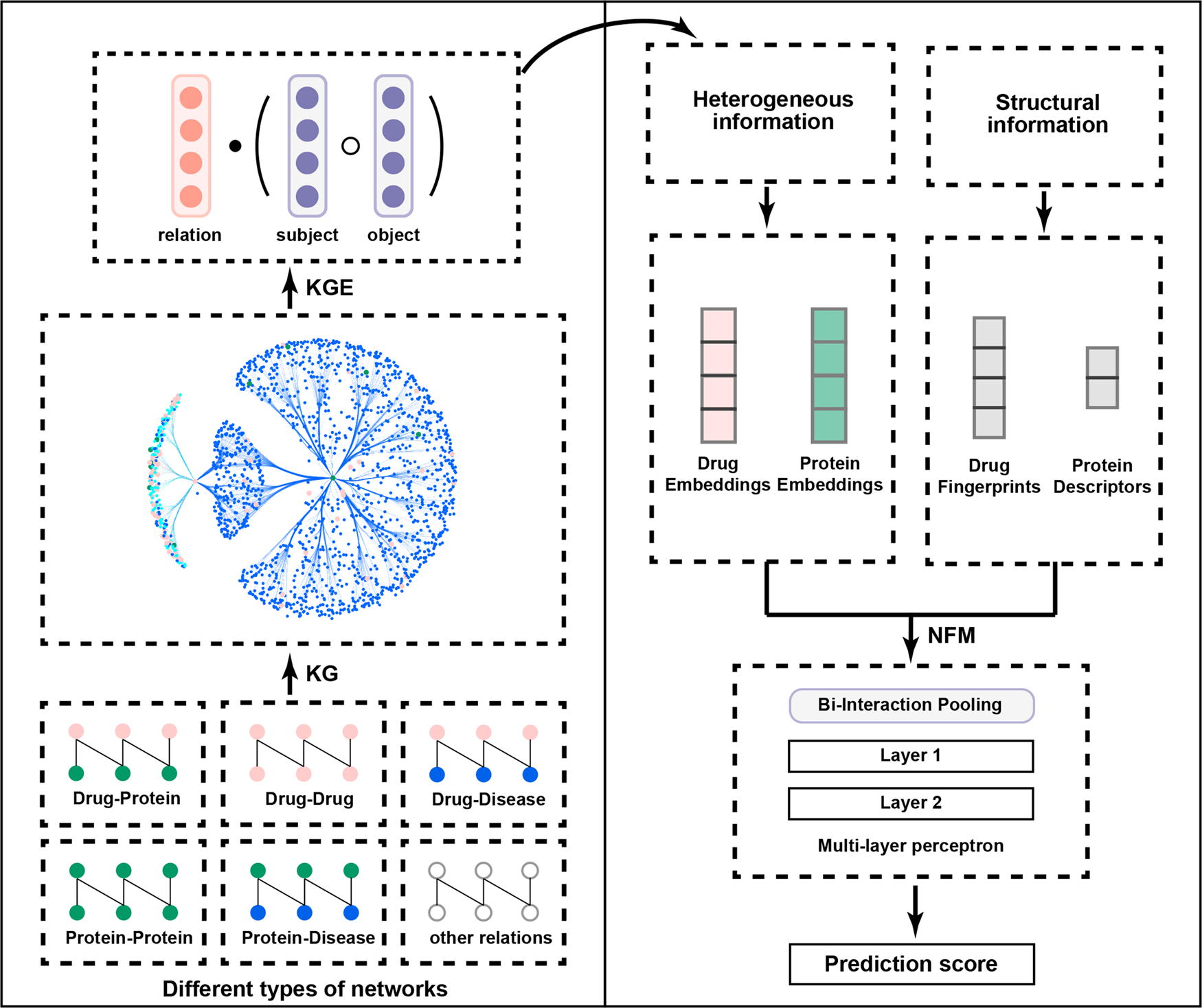
A unified drug–target interaction prediction framework based on knowledge graph and recommendation system | Nature Communications
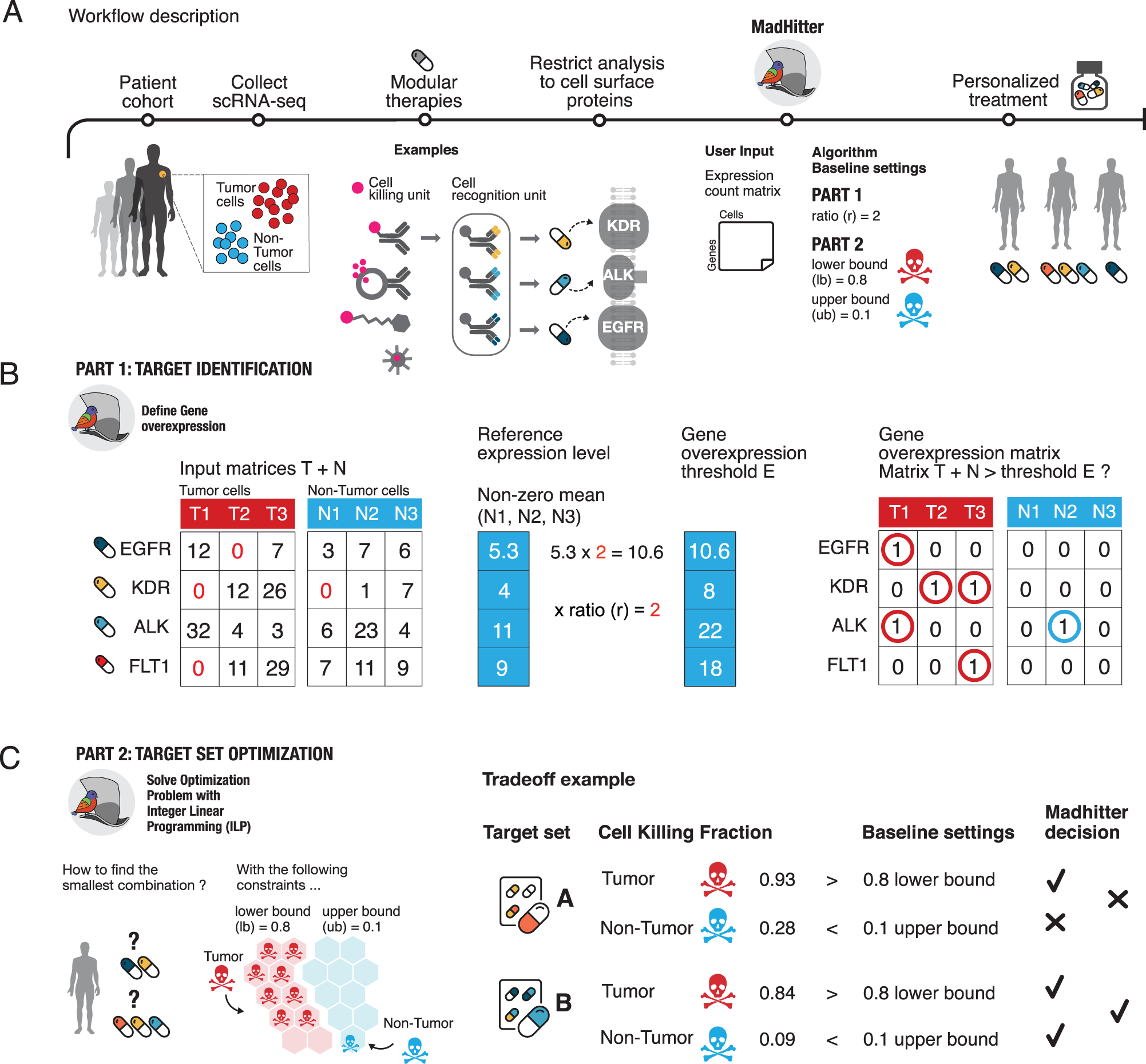
The landscape of receptor-mediated precision cancer combination therapy via a single-cell perspective | Nature Communications